Researchers Uncover Genetic Insights Into Human Diseases Through ORIP’s National Primate Research Centers
Nonhuman primates (NHPs) offer valuable insights into many diseases that affect human health because they are genetically and physiologically very similar to humans (Figure 1). ORIP’s National Primate Research Centers (NPRC) Consortium maintains NHP colonies for use by the biomedical research community. Dr. Jeffrey Rogers, Core Scientist at the Wisconsin NPRC and Associate Professor at the Baylor College of Medicine (BCM), is leading efforts to identify naturally occurring mutations within NHP colonies across the NPRCs and other primate centers. By identifying naturally occurring mutations and applying machine-learning approaches, researchers can discover and develop animal models more quickly and efficiently than is possible through other methods, such as CRISPR gene editing. This work will help researchers study NHP models for both rare and common chronic diseases, leading to potential new treatments for patients.

Dr. Rogers emphasized that although human genome sequencing has enabled significant progress in the field of genetic disorders, NHPs can provide a missing link for understanding the mutations that contribute to diseases in humans. He explained that the NHP colonies encompass a wealth of diverse genetic data. “Many people assume that rhesus macaques in research colonies are like laboratory mice, that they’re inbred and don’t have much genetic variation. But rhesus macaques have more than twice as much genetic variation as humans,” he explained. “If you sequence 1,000 macaques, you’re going to find more genetic variation than you would by sequencing an equivalent number of people.”
The idea for a specific project on the genetics of visual system disorders was sparked by an anecdotal observation by Dr. Rogers’ colleague, Dr. Jeffrey Roberts. Dr. Roberts, Associate Director of Primate Services at the California NPRC, observed that several animals in his Center appeared to have visual impairments and were shielding their eyes from direct sunlight in ways not usually observed in other NHPs. Dr. Sara Thomasy and Dr. Ala Moshiri (both affiliated with the California NPRC) examined the animals and found that they had severe dysfunction of cone photoreceptors in their retinas. After sequencing DNA samples from the affected individuals, the team discovered that the animals had a homozygous mutation in a gene that is known to cause achromatopsia (an inherited form of color blindness) in humans.1 This finding led to a new gene therapy for achromatopsia that has been successfully tested in naturally affected rhesus macaques by Dr. Timothy Stout (BCM) and Drs. Thomasy and Moshiri. Their work may lead to new treatments in humans. NHPs are well suited for this research because their eyes have a macula (i.e., the part of the eye that is required for seeing color and fine details) that is similar to humans. Other animals—such as rodents, cats, and dogs—do not possess this structure.
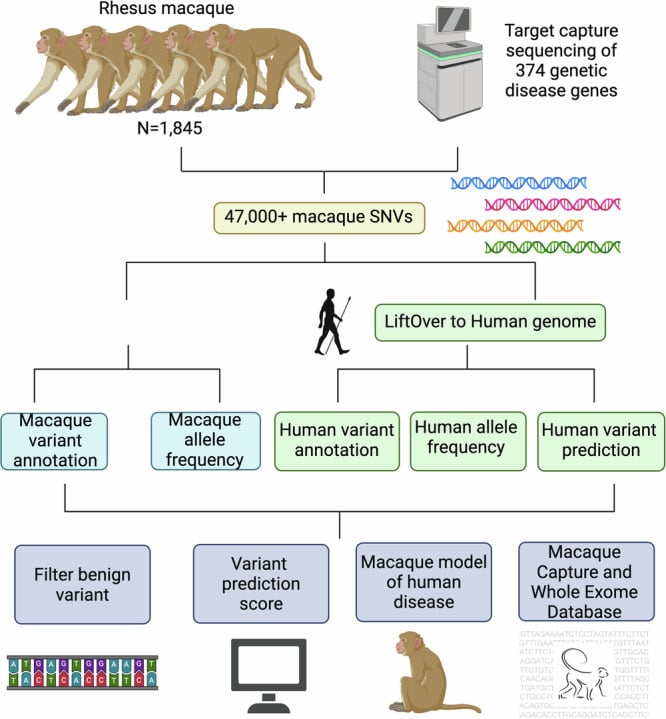
To expand on this finding, the collaborative team—which included Dr. Rui Chen, Professor, and Dr. Jun Wang, Postdoctoral Associate (both affiliated with BCM)—received support from the National Eye Institute (U24EY029904) to identify new models for human retinal diseases. The team conducted a survey of genetic variation among rhesus macaques housed at the primate centers. From this analysis, the team identified genetic variations in the 374 genes that occurred at varying frequencies in the rhesus macaque and human populations (Figure 2).
By integrating the genetic variant frequencies from the two species with established in silico prediction scores, Dr. Wang developed machine-learning models to predict the pathogenicity of missense variants. These models were trained using the annotated genetic variants from the ClinVar database. Among the seven machine learning models developed, the random forest model performed best at recognizing known pathogenic mutations, and better than the individual prediction scores it incorporated. Dr. Wang then used the random forest model to make novel predictions regarding the effects of the identified macaque variants; these results are now available to the research community. In addition, the team also identified animals with a mutation in the OPA1 gene, which is associated with autosomal dominant optic atrophy (a progressive eye disease) in humans.2 Subsequent analysis showed that the animals carrying the OPA1 genetic mutations show visual system pathology very much like humans with damaging OPA1 mutations.
By studying natural mutations in animal models, Dr. Rogers and his colleagues have gained insight into several other human disorders, including retinal diseases and blindness, cancer, polycystic ovarian syndrome, and endometriosis. This approach is also useful for characterizing complex structural variants, which are difficult to detect accurately. The information from these studies will be available to the research community through a public database. Investigators will be able to search the database for NHP mutations that have been linked to diseases in humans.
Dr. Rogers underscored the importance of data sharing and discussions across multiple disciplines to further research on this topic. “There is a tremendous opportunity for new collaborations—interactions between experts in primate models and experts in various human genetic diseases,” he reflected. Dr. Rogers also highlighted the value of ORIP’s support for the NPRCs, which helped make this work possible. “We would not have been able to do this work without the funding provided to generate the pedigrees and the background information for the primate centers,” he stated. “The NPRC program is essential—it is tremendously important.”
Future sequencing studies will be crucial to uncovering even more NHP genetic variants that are associated with diseases. By expanding the research to include other NHP species—such as marmosets, baboons, and squirrel monkeys—researchers can identify more naturally occurring mutations within the NPRCs. “Substantial numbers of new research opportunities are already present in the colonies that, unfortunately, we’re not able to take advantage of because we haven’t identified the mutations,” Dr. Rogers emphasized. “I believe that dozens of naturally occurring models are available in the primate centers today that we haven’t yet discovered.”
The primary goal of the NPRC program is to facilitate the effective use of NHPs by scientists engaged in biomedical research. The NPRC program complements and enables the missions of NIH institutes and centers (ICs) by providing the animals, facilities, expertise, and resources required to enable NHP research in specific disease areas. ORIP’s Division of Comparative Medicine funds seven NPRCs, which are centralized facilities in various parts of the country and are available to investigators nationwide. The majority of researchers who use the NPRC physical and intellectual infrastructure are funded by the U.S. Public Health Service, particularly NIH ICs. More information on the NPRC program can be found through ORIP’s Research Resources Directory.
References
1 Moshiri A, Chen R, Kim S, et al. A nonhuman primate model of inherited retinal disease. J Clin Invest. 2019 Jan 22;129(2):863–874. doi:10.1172/JCI123980.
2 Wang J, Wang M, Moshiri A, et al. Genetic diversity of 1,845 rhesus macaques improves genetic variation interpretation and identifies disease models. Nat Commun. 2024 Jul 5;15(1):5658. doi:10.1038/s41467-024-49922-6.



