S10 Instrumentation Programs Enhance Capabilities for Proteomics Research at Cleveland Clinic’s Lerner Research Institute
Understanding the molecular composition and structure of proteins is essential to the mechanistic study of human health and diseases and requires the use of state-of-the-art scientific instruments. Two recent awards to Cleveland Clinic’s Lerner Research Institute (LRI) through ORIP’s S10 Shared Instrumentation Programs (S10OD023436 and S10OD030398) have enhanced the organization’s capacity for proteomics, the study of protein structure and function in different biological contexts. These awards funded the acquisition of two high-resolution mass spectrometers (MS)—a Thermo Scientific Fusion Lumos MS and a Bruker trapped ion mobility spectrometry quadrupole time of flight (QTOF) MS—for LRI’s Proteomics and Metabolomics Core, led by Dr. Belinda Willard, Director of LRI’s Proteomics and Metabolomics Core (Figure 1).

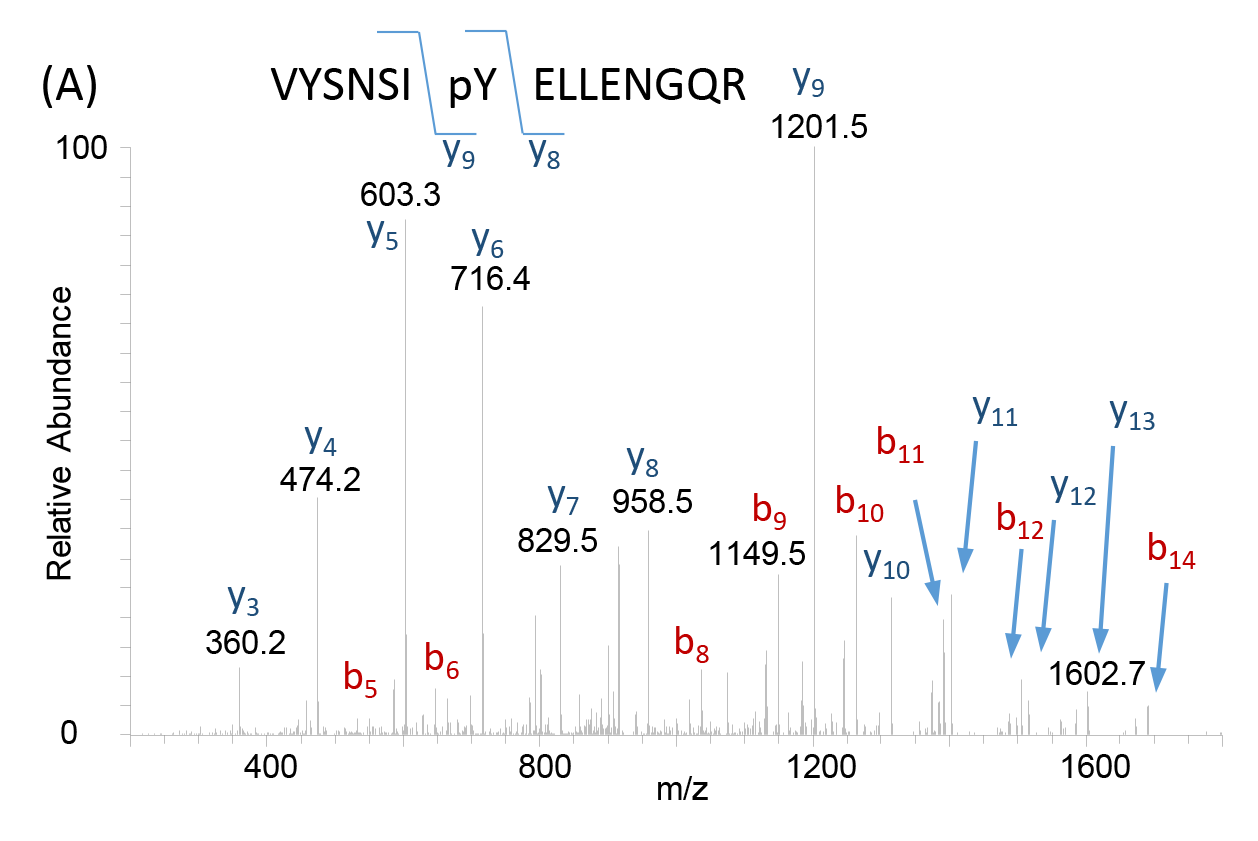
Following ionization of the biological sample, an MS produces a spectrum or a plot of the ion signal pattern where each ion has a specific mass-to-charge ratio, allowing analysis of elemental chemical characteristics and determination of chemical and structural signatures in the sample. These data are used for identification of unknown compounds, quantification of known compounds, and delineation of structural properties for molecules of interest (Figure 2). As a critical tool in proteomics, MS serve as a valuable resource in many fields of biomedicine, enabling molecular discoveries and development of new therapeutics for disease.
Dr. Willard oversees all projects conducted using the two S10-funded instruments. She plays a key role in managing the operation of the instruments, implementation of institutional prioritization, and formulation of experimental design, data acquisition, and data analysis. The S10-funded instruments have provided the facility with crucial capabilities for research. The core instrumentation runs continuously, collecting data from various biological samples (e.g., tissue samples, cell-derived samples, clinical samples) provided by more than 100 laboratories. Dr. Willard noted that investigators use MS across a broad array of research areas, including the study of changes in protein abundance and identification of novel substrates for proteases associated with chronic diseases. The Lumos MS has contributed directly to several major research outcomes.
Dr. Srinivasan Dasarathy, a Professor in the Department of Molecular Medicine at the Case Western Reserve University (CWRU) School of Medicine, is one of a dozen users of LRI’s Lumos MS. Dr. Dasarathy studies molecular changes in skeletal muscle and myotubes under conditions of hyperammonemia, a disorder that is present in such diseases as liver cirrhosis, heart failure, and chronic obstructive pulmonary disease. His group is working to characterize the proteomic response to hyperammonemia and ethanol. In a series of experiments, protein-level data were matched to changes in chromosomal conformation and transcriptomics, and these experiments identified a novel regulatory pathway that plays a role in how cells respond to hyperammonemia. These experiments are an example of a systems biology approach—an approach that holistically analyzes tissues and cells, with the goal of identifying targets for the development of novel disease-modifying therapeutics.1
Dr. Christopher Hine, an Assistant Professor in the Department of Molecular Medicine at the CWRU School of Medicine, has used LRI’s Lumos MS to study shifts in post-translational protein persulfidation (i.e., addition of sulfur molecules) associated with dietary restriction2 and glioblastoma cancer.3 In one study, Dr. Hine found that short-term caloric restriction in young adult mice—as well as long term intermittent fasting in aged mice—led to the enrichment of protein persulfidation in the liver, kidney, skeletal muscle, and brain and a decreased protein persulfidation in the heart. Protein persulfidation enrichment, a function related to the protein’s and cellular defense responses to oxidative injury, was associated with a multitude of cellular metabolic pathways in these tissues. In a second study, Dr. Hine—in collaboration with Dr. Justin Lathia, Melvin H. Burkhardt Endowed Chair for Neuro-Oncology Clinical Research and Staff at LRI and Professor in the Department of Molecular Medicine at the CWRU School of Medicine—found that a high-fat diet decreased protein persulfidation in the glioma-containing brains of mice. In human glioblastoma patients, the team found a decrease of protein persulfidation in tumor biopsies compared to non-cancerous biopsies, ultimately helping them conclude that protein persulfidation and its gaseous precursor hydrogen sulfide act as tumor suppressors against glioblastoma.
Dr. Willard emphasized that the capabilities provided by the high-resolution MS instruments meet a growing need within the scientific community. The instruments have expanded the core facility’s services, which now include quantitation of protein abundances using label-free and isobaric labeling approaches. Dr. Willard explained that the QTOF MS, which was installed in August 2021, provides higher-throughput capabilities for researchers, processing samples in 30 minutes. The Lumos MS, which was acquired in 2017, processes samples in 2 hours.
“The Shared Instrumentation mechanism has fostered the LRI Proteomics and Metabolomics Core to service investigators at a higher level,” says Dr. Willard. “This funding mechanism has been crucial in expanding the capabilities of the core to quantify proteins at a depth and accuracy that was not possible with our previous equipment.” Dr. Willard said that the core facility is available to all investigators representing different fields of research from across the scientific community. The user base consists of investigators from Cleveland Clinic, as well as CWRU, Cleveland State University, Kent State University, Lake Erie College of Osteopathic Medicine, Michigan State University, and Northeast Ohio Medical University.
“We have an open policy, and we’re happy to help researchers with their experiments,” Dr. Willard says. “We make the resource available to anyone who comes to our core and has an interest in working with us.”
For more information on ORIP’s S10 Shared Instrumentation Programs, please visit orip.nih.gov/construction-and-instruments/s10-instrumentation-programs.
References
1 Welch N, Singh SS, Kumar A, et al. Integrated multiomics analysis identifies molecular landscape perturbations during hyperammonemia in skeletal muscle and myotubes. J Biol Chem. 2021;297(3):101023. doi:10.1016/j.jbc.2021.101023.
2 Bithi N, Link C, Henderson YO, et al. Dietary restriction transforms the mammalian protein persulfidome in a tissue-specific and cystathionine γ-lyase-dependent manner. Nat Commun. 2021;12:1745. doi:10.1038/s41467-021-22001-w.
3 Silver DJ, Roversi GA, Bithi N, et al. Severe consequences of a high-lipid diet include hydrogen sulfide dysfunction and enhanced aggression in glioblastoma. J Clin Invest. 2021;131(17):e138276. doi:10.1172/JCI138276.



