A Drug Discovery Platform To Rapidly Diagnose Patients
Precision disease modeling has gained increased interest in the research community in recent years and remains at the forefront of personalized medicine approaches for developing models of human disease. Informed by recommendations1 from the 2013 “Animal Models and Personalized Medicine Symposium” to coordinate research projects to facilitate knowledge and resource sharing, the Office of Research Infrastructure Programs (ORIP) at the National Institutes of Health (NIH) launched the Precision Disease Modeling Initiative in 2014. Subsequently, in July 2015, ORIP funded three Pilot Disease Modeling Centers (U54). The Mount Sinai Pilot Center is one of the three centers established to develop demonstration pipelines for preclinical scientific discovery and disease modeling. In a multidisciplinary approach, researchers collaborate to link advances in animal genomics and technology with personalized medicine efforts in human subjects.

Dr. Ross L. Cagan, principal investigator at the Mount Sinai Pilot Center for Precision Disease Modeling and professor at the Icahn School of Medicine at Mount Sinai, began his ORIP U54 project—A New Disease Platform Leveraging Complex Drosophila and Mammalian Models—by assembling a multidisciplinary research team to develop a platform that seamlessly moves from (1) sequencing to (2) Drosophila transgenics to (3) human disease models. Subsequently, he has built a unique pipeline (i.e., functional network platform) designed to identify treatments for colorectal cancer and other human diseases.
Dr. Cagan uses fruit flies (Drosophila), mouse genetics, and stem cell technology to build his personalized drug discovery platform. Drosophila, the primary model used in developmental genetic studies, allows researchers to analyze both the tumor and its immediate surrounding environment. The fruit fly’s short life span, rapid reproduction cycle, simple genetic profile, and anatomic features make it an ideal model organism2 (Figure 1).
An additional advantage of using the fruit fly as a model for drug discovery is the ability to modify a large number of genes in a single organism, which is useful for screening a sizeable number of drugs (single agents or combinations) in the whole animal. Dr. Cagan and his laboratory, along with Icahn School of Medicine collaborators Drs. Arvin Dar and Avner Schlessinger and their teams, have designed a unique platform that combines complex genetic fruit fly models (i.e., fly avatars) with a high-throughput multidrug-screening approach to identify novel treatments (Figure 2).
It is noteworthy that Dr. Cagan and his colleagues’ personalized medicine approach3 has informed clinical trials at the Mount Sinai Center for Personalized Cancer Therapeutics (CPCT) (Figure 3).
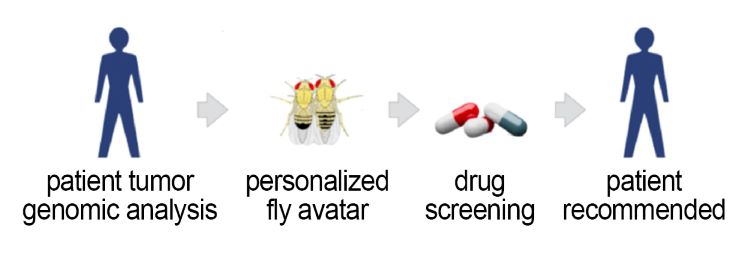
First, a comprehensive genomic analysis of a cancer patient’s tumor and normal DNA is performed. Second, using the pipeline, a personalized fruit fly avatar captures a portion of the patient’s tumor’s genomic complexity (typically eight to 15 altered genes) and cell abnormalities. Next, a high-throughput screening of patient-derived avatar flies is performed against a library of 1,500 drugs approved by the U.S. Food and Drug Administration. The selected drugs are mixed into the insects’ food, and the scientists track their survival. Scientific and pharmacy tumor boards review the data, and a personalized treatment plan is developed and approved by the Mount Sinai Internal Review Board. The patient then is treated with a novel two- to three-drug combination cocktail.
Dr. Cagan and his colleagues are glad to report that their novel platform affords better therapeutic efficacy and lower whole-animal toxicity and now is validated for the treatment of colorectal3 and thyroid cancers,4 RASopathies, and tauopathies. This novel treatment approach also has improved outcomes for cancer patients experiencing relapse after previous therapies.3,5,6 For example, one study revealed that a patient with colorectal cancer who failed two previous lines of treatment strongly responded to a combination of a MEK inhibitor (trametinib) and a bisphosphonate (zoledronate) based on personalized recommendations generated from this platform, with a partial response of 46 percent tumor shrinkage over 11 months.3 This research shows great promise in achieving the goal of personalized medicine by treating each patient with the most beneficial and effective drugs with minimal side effects. Overall, this platform is cost-effective, rapid, and disease-agnostic and enables a simple readout for efficacy and toxicity.
Exploring the relationship between genomic complexity and drug resistance has enabled Dr. Cagan’s work to support a fly-to-bedside clinical trial led by the CPCT. To date, the trial has enrolled 29 patients, made 23 drug recommendations, and treated nine patients. In an integrated approach, Dr. Cagan and colleagues have combined the Drosophila model with structural prediction, virtual screening, and computational modeling, thereby creating a unique personal drug discovery platform.7 In collaboration with Dr. Laura Towart, founder and CEO of London-based startup My Personal Therapeutics, Dr. Cagan has taken this personal discovery process technology and personalized fruit fly avatars to market.
The Mount Sinai Pilot Center for Precision Disease Modeling research teams generated a broad array of technologies and resources, including Drosophila models now being distributed through the Bloomington Drosophila Stock Center (another ORIP supported resource). They also generated powerful genetic tools, such as the vectors to create up-to-18+-hit cancer models that capture a level of complexity in a transgenic system not seen elsewhere. Furthermore, a variety of drug or chemical compound computational platforms and a set of patient-derived xenograft mouse models of colorectal cancer were generated and tested.
Dr. Cagan and his colleagues clearly demonstrate how ORIP funding facilitates bench-to-bedside discoveries, with the hope of improving outcomes for patients. Dr. Cagan stated that, “I am a geneticist who uses a broad palette of tools to explore disease. … I have mentored approximately 40 students and postdoctoral fellows who are now doing amazing science around the planet.” In May 2020, the Cagan laboratory relocated to the University of Glasgow, where they will continue to expand the application of this platform developed with support from ORIP’s Precision Disease Modeling Initiative.
References
- ORIP. 2013. “Animal models and personalized medicine.” dpcpsi.nih.gov/sites/default/files/Animal_Models_and_Personalized_Medicine_Meeting_Summary.pdf
- Cagan RL, Zon LI, White RM. Modeling cancer with flies and fish. Developmental Cell 2019;49 (3):317–24.
- Bangi E, Ang C, Smibert P, Uzilov AV, Teague AG, Antipin Y, et al. A personalized platform identifies trametinib plus zoledronate for a patient with KRAS-mutant metastatic colorectal cancer. Science Advances 2019;22;5 (5):eaav6528.
- Das TK, Cagan RL. A Drosophila approach to thyroid cancer therapeutics. Drug Discovery Today: Technology 2013;10 (1):e65–71.
- Sonoshita M, Cagan RL. Modeling human cancers in Drosophila. Current Topics in Developmental Biology 2017;121:287–309
- Ung PMU, Sonoshita M, Scopton AP, Dar AC, Cagan RL, Schlessinger A. Integrated computational and Drosophila cancer model platform captures previously unappreciated chemicals perturbing a kinase network. PLoS Computational Biology 2019;15 (4):e1006878.
- Sonoshita M, Scopton AP, Ung PMU, Murray MA, Silber L, Maldonado AY, et al. A whole-animal platform to advance a clinical kinase inhibitor into new disease space. Nature Chemical Biology 2018;14(3):291–298.



